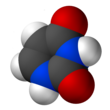
Uracil (/ˈjʊərəsɪl/) (symbol U or Ura) is one of the four nucleobases in the nucleic acid RNA. The others are adenine (A), cytosine (C), and guanine (G). In RNA, uracil binds to adenine via two hydrogen bonds. In DNA, the uracil nucleobase is replaced by thymine (T). Uracil is a demethylated form of thymine.

| |||
| |||
| Names | |||
|---|---|---|---|
| Preferred IUPAC name
Pyrimidine-2,4(1H,3H)-dione | |||
Other names
| |||
| Identifiers | |||
3D model (JSmol)
|
| ||
| 3DMet | |||
| 606623 | |||
| ChEBI | |||
| ChEMBL | |||
| ChemSpider | |||
| DrugBank | |||
| ECHA InfoCard | 100.000.565 | ||
| EC Number |
| ||
| 2896 | |||
| KEGG | |||
PubChem CID
|
|||
| RTECS number |
| ||
| UNII | |||
CompTox Dashboard (EPA)
|
|||
| |||
| Properties | |||
| C4H4N2O2 | |||
| Molar mass | 112.08676 g/mol | ||
| Appearance | Solid | ||
| Density | 1.32 g/cm3 | ||
| Melting point | 335 °C (635 °F; 608 K)[1] | ||
| Boiling point | N/A – decomposes | ||
| Soluble | |||
| Hazards | |||
| Occupational safety and health (OHS/OSH): | |||
Main hazards
|
carcinogen and teratogen with chronic exposure | ||
| GHS labelling: | |||
 
| |||
| Warning | |||
| H315, H319, H335, H361 | |||
| P201, P202, P261, P264, P271, P280, P281, P302+P352, P304+P340, P305+P351+P338, P308+P313, P312, P321, P332+P313, P337+P313, P362, P403+P233, P405, P501 | |||
| NFPA 704 (fire diamond) | |||
| Flash point | Non-flammable | ||
| Related compounds | |||
Related compounds
|
Thymine Cytosine | ||
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
| |||
Uracil is a common and naturally occurring pyrimidine derivative.[2] The name "uracil" was coined in 1885 by the German chemist Robert Behrend, who was attempting to synthesize derivatives of uric acid.[3] Originally discovered in 1900 by Alberto Ascoli, it was isolated by hydrolysis of yeast nuclein;[4] it was also found in bovine thymus and spleen, herring sperm, and wheat germ.[5] It is a planar, unsaturated compound that has the ability to absorb light.[6]
Uracil that was formed extraterrestrially has been detected in the Murchison meteorite,[7] in a near-Earth asteroid,[8] and possibly on the surface of the moon Titan.[9] It has been synthesized under cold laboratory conditions similar to outer space, from pyrimidine embedded in water ice and exposed to ultraviolet light.[10]
Properties edit
In RNA, uracil base-pairs with adenine and replaces thymine during DNA transcription. Methylation of uracil produces thymine.[11] In DNA, the evolutionary substitution of thymine for uracil may have increased DNA stability and improved the efficiency of DNA replication (discussed below). Uracil pairs with adenine through hydrogen bonding. When base pairing with adenine, uracil acts as both a hydrogen bond acceptor and a hydrogen bond donor. In RNA, uracil binds with a ribose sugar to form the ribonucleoside uridine. When a phosphate attaches to uridine, uridine 5′-monophosphate is produced.[6]
Uracil undergoes amide-imidic acid tautomeric shifts because any nuclear instability the molecule may have from the lack of formal aromaticity is compensated by the cyclic-amidic stability.[5] The amide tautomer is referred to as the lactam structure, while the imidic acid tautomer is referred to as the lactim structure. These tautomeric forms are predominant at pH 7. The lactam structure is the most common form of uracil.
Uracil also recycles itself to form nucleotides by undergoing a series of phosphoribosyltransferase reactions.[2] Degradation of uracil produces the substrates β-alanine, carbon dioxide, and ammonia.[2]
- C4H4N2O2→ H3NCH2CH2COO− + NH+4 + CO2
Oxidative degradation of uracil produces urea and maleic acid in the presence of H2O2 and Fe2+ or in the presence of diatomic oxygen and Fe2+.
Uracil is a weak acid. The first site of ionization of uracil is not known.[12] The negative charge is placed on the oxygen anion and produces a pKa of less than or equal to 12. The basic pKa = −3.4, while the acidic pKa = 9.389. In the gas phase, uracil has four sites that are more acidic than water.[13]
In DNA edit
Uracil is rarely found in DNA, and this may have been an evolutionary change to increase genetic stability. This is because cytosine can deaminate spontaneously to produce uracil through hydrolytic deamination. Therefore, if there were an organism that used uracil in its DNA, the deamination of cytosine (which undergoes base pairing with guanine) would lead to formation of uracil (which would base pair with adenine) during DNA synthesis. Uracil-DNA glycosylase excises uracil bases from double-stranded DNA. This enzyme would therefore recognize and cut out both types of uracil – the one incorporated naturally, and the one formed due to cytosine deamination, which would trigger unnecessary and inappropriate repair processes.[14]
This problem is believed to have been solved in terms of evolution, that is by "tagging" (methylating) uracil. Methylated uracil is identical to thymine. Hence the hypothesis that, over time, thymine became standard in DNA instead of uracil. So cells continue to use uracil in RNA, and not in DNA, because RNA is shorter-lived than DNA, and any potential uracil-related errors do not lead to lasting damage. Apparently, either there was no evolutionary pressure to replace uracil in RNA with the more complex thymine, or uracil has some chemical property that is useful in RNA, which thymine lacks. Uracil-containing DNA still exists, for example in
- DNA of several phages[15]
- Endopterygote development
- Hypermutations during the synthesis of vertebrate antibodies.[citation needed]
Synthesis edit
Biological edit
Organisms synthesize uracil, in the form of uridine monophosphate (UMP), by decarboxylating orotidine 5'-monophosphate (orotidylic acid). In humans this decarboxylation is achieved by the enzyme UMP synthase. In contrast to the purine nucleotides, the pyrimidine ring (orotidylic acid) that leads uracil is synthesized first and then linked to ribose phosphate, forming UMP.[16]
Laboratory edit
There are many laboratory synthesis of uracil available. The first reaction is the simplest of the syntheses, by adding water to cytosine to produce uracil and ammonia:[2]
- C4H5N3O + H2O → C4H4N2O2 + NH3
The most common way to synthesize uracil is by the condensation of malic acid with urea in fuming sulfuric acid:[5]
- C4H4O4 + NH2CONH2 → C4H4N2O2 + 2 H2O + CO
Uracil can also be synthesized by a double decomposition of thiouracil in aqueous chloroacetic acid.[5]
Photodehydrogenation of 5,6-diuracil, which is synthesized by beta-alanine reacting with urea, produces uracil.[17]
Prebiotic edit
In 2009, NASA scientists reported having produced uracil from pyrimidine and water ice by exposing it to ultraviolet light under space-like conditions.[10] This suggests a possible natural original source for uracil.[18] In 2014, NASA scientists reported that additional complex DNA and RNA organic compounds of life, including uracil, cytosine and thymine, have been formed in the laboratory under outer space conditions, starting with ice, pyrimidine, ammonia, and methanol, which are compounds found in astrophysical environments.[19] Pyrimidine, like polycyclic aromatic hydrocarbons (PAHs), a carbon-rich chemical found in the Universe, may have been formed in red giants or in interstellar dust and gas clouds.[20]
Based on 12C/13C isotopic ratios of organic compounds found in the Murchison meteorite, it is believed that uracil, xanthine, and related molecules can also be formed extraterrestrially.[7] Data from the Cassini mission, orbiting in the Saturn system, suggests that uracil is present in the surface of the moon Titan.[9] In 2023, uracil was observed in a sample from 162173 Ryugu, a near-Earth asteroid, with no exposure to Earth's biosphere, giving further evidence for synthesis in space.[8]
Reactions edit
Uracil readily undergoes regular reactions including oxidation, nitration, and alkylation. While in the presence of phenol (PhOH) and sodium hypochlorite (NaOCl), uracil can be visualized in ultraviolet light.[5] Uracil also has the capability to react with elemental halogens because of the presence of more than one strongly electron donating group.[5]
Uracil readily undergoes addition to ribose sugars and phosphates to partake in synthesis and further reactions in the body. Uracil becomes uridine, uridine monophosphate (UMP), uridine diphosphate (UDP), uridine triphosphate (UTP), and uridine diphosphate glucose (UDP-glucose). Each one of these molecules is synthesized in the body and has specific functions.
When uracil reacts with anhydrous hydrazine, a first-order kinetic reaction occurs and the uracil ring opens up.[21] If the pH of the reaction increases to > 10.5, the uracil anion forms, making the reaction go much more slowly. The same slowing of the reaction occurs if the pH decreases, because of the protonation of the hydrazine.[21] The reactivity of uracil remains unchanged, even if the temperature changes.[21]
Uses edit
Uracil's use in the body is to help carry out the synthesis of many enzymes necessary for cell function through bonding with riboses and phosphates.[2] Uracil serves as allosteric regulator and coenzyme for reactions in animals and in plants.[22] UMP controls the activity of carbamoyl phosphate synthetase and aspartate transcarbamoylase in plants, while UDP and UTP regulate CPSase II activity in animals. UDP-glucose regulates the conversion of glucose to galactose in the liver and other tissues in the process of carbohydrate metabolism.[22] Uracil is also involved in the biosynthesis of polysaccharides and the transportation of sugars containing aldehydes.[22] Uracil is important for the detoxification of many carcinogens, for instance those found in tobacco smoke.[23] Uracil is also required to detoxify many drugs such as cannabinoids (THC)[24] and morphine (opioids).[25] It can also slightly increase the risk for cancer in unusual cases in which the body is extremely deficient in folate.[26] The deficiency in folate leads to increased ratio of deoxyuridine monophosphates (dUMP)/deoxythymidine monophosphates (dTMP) and uracil misincorporation into DNA and eventually low production of DNA.[26]
Uracil can be used for drug delivery and as a pharmaceutical. When elemental fluorine reacts with uracil, they produce 5-fluorouracil. 5-Fluorouracil is an anticancer drug (antimetabolite) used to masquerade as uracil during the nucleic acid replication process.[2] Because 5-fluorouracil is similar in shape to, but does not undergo the same chemistry as, uracil, the drug inhibits RNA transcription enzymes, thereby blocking RNA synthesis and stopping the growth of cancerous cells.[2] Uracil can also be used in the synthesis of caffeine.[27] Uracil has also shown potential as a HIV viral capsid inhibitor.[28] Uracil derivatives have antiviral, anti-tubercular and anti-leishmanial activity.[29][30][31]
Uracil can be used to determine microbial contamination of tomatoes. The presence of uracil indicates lactic acid bacteria contamination of the fruit.[32] Uracil derivatives containing a diazine ring are used in pesticides.[33] Uracil derivatives are more often used as antiphotosynthetic herbicides, destroying weeds in cotton, sugar beet, turnips, soya, peas, sunflower crops, vineyards, berry plantations, and orchards.[33] Uracil derivatives can enhance the activity of antimicrobial polysaccharides such as chitosan.[34]
In yeast, uracil concentrations are inversely proportional to uracil permease.[35]
Mixtures containing uracil are also commonly used to test reversed-phase HPLC columns. As uracil is essentially unretained by the non-polar stationary phase, this can be used to determine the dwell time (and subsequently dwell volume, given a known flow rate) of the system.
References edit
- ^ Myers RL (2007). "Chapter 29: Cytosine Thymine and Uracil". The 100 most important chemical compounds : a reference guide. Westport, Conn.: Greenwood Press. pp. 92–93. ISBN 978-0-313-33758-1.
- ^ a b c d e f g Garrett RH, Grisham CM (1997). Principles of Biochemistry with a Human Focus. United States: Brooks/Cole Thomson Learning.
- ^ Behrend R (1885). "Versuche zur Synthese von Körpern der Harnsäurereihe" [Experiments on the synthesis of substances in the uric acid series]. Annalen der Chemie. 229 (1–2): 1–44. doi:10.1002/jlac.18852290102.
Dasselbe stellt sich sonach als Methylderivat der Verbindung: welche ich willkürlich mit dem Namen Uracil belege, dar.
[The same compound is therefore represented as the methyl derivative of the compound, which I will arbitrarily endow with the name ‘uracil’.] - ^ Ascoli A (1900). "Über ein neues Spaltungsprodukt des Hefenucleins" [On a new cleavage product of nucleic acid from yeast]. Zeitschrift für Physiologische Chemie. 31 (1–2): 161–164. doi:10.1515/bchm2.1901.31.1-2.161. Archived from the original on 12 May 2018.
- ^ a b c d e f Brown DJ, Evans RF, Cowden WB, Fenn MD (1994). Taylor EC (ed.). The Pyrimidines. Heterocyclic Compounds. Vol. 52. New York, NY: Wiley. ISBN 9780471506560. Archived from the original on 12 May 2018.
- ^ a b Horton HR, Moran LA, Ochs RS, Rawn DJ, Scrimgeour KG (2002). Principles of Biochemistry (3rd ed.). Upper Saddle River, NJ: Prentice Hall. ISBN 9780130266729.
- ^ a b Martins Z, Botta O, Fogel ML, Sephton MA, Glavin DP, Watson JS, et al. (2008). "Extraterrestrial nucleobases in the Murchison meteorite". Earth and Planetary Science Letters. 270 (1–2): 130–136. arXiv:0806.2286. Bibcode:2008E&PSL.270..130M. doi:10.1016/j.epsl.2008.03.026. S2CID 14309508.
- ^ a b Oba Y, Koga T, Takano Y, Ogawa NO, Ohkouchi N, Sasaki K, et al. (2023). "Uracil in the carbonaceous asteroid (162173) Ryugu". Nature Communications. 14 (1): 1292. Bibcode:2023NatCo..14.1292O. doi:10.1038/s41467-023-36904-3. PMC 10030641. PMID 36944653.
- ^ a b Clark RN, Pearson N, Brown RH, Cruikshank DP, Barnes J, Jaumann R, et al. (2012). "The Surface Composition of Titan". American Astronomical Society. 44: 201.02. Bibcode:2012DPS....4420102C.
- ^ a b Nuevo, Michel; Milam, Stefanie N.; Sandford, Scott A.; Elsila, Jamie E.; Dworkin, Jason P. (2009). "Formation of Uracil from the Ultraviolet Photo-Irradiation of Pyrimidine in Pure H2O Ices". Astrobiology. 9 (7): 683–695. Bibcode:2009AsBio...9..683N. doi:10.1089/ast.2008.0324. ISSN 1531-1074. PMID 19778279.
- ^ "MadSciNet: The 24-hour exploding laboratory". www.madsci.org. Archived from the original on 18 July 2005.
- ^ Zorbach WW, Tipson RS (1973). Synthetic Procedures in Nucleic Acid Chemistry: Physical and physicochemical aids in determination of structure. Vol. 2. New York, NY: Wiley-Interscience. ISBN 9780471984184.
- ^ Kurinovich MA, Lee JK (August 2002). "The acidity of uracil and uracil analogs in the gas phase: four surprisingly acidic sites and biological implications". Journal of the American Society for Mass Spectrometry. 13 (8): 985–995. doi:10.1016/S1044-0305(02)00410-5. PMID 12216739.
- ^ Békési A, Vértessy BG (2011). "Uracil in DNA: error or signal?". Science in School: 18. Archived from the original on 23 March 2016.
- ^ Wang Z, Mosbaugh DW (March 1988). "Uracil-DNA glycosylase inhibitor of bacteriophage PBS2: cloning and effects of expression of the inhibitor gene in Escherichia coli". Journal of Bacteriology. 170 (3): 1082–1091. doi:10.1128/JB.170.3.1082-1091.1988. PMC 210877. PMID 2963806.
- ^ Löffler, Monika; Zameitat, Elke (2004). "Pyrimidine Biosynthesis". Encyclopedia of Biological Chemistry. Elsevier. pp. 600–605. doi:10.1016/b0-12-443710-9/00574-3. ISBN 9780124437104.
- ^ Chittenden GJ, Schwartz AW (September 1976). "Possible pathway for prebiotic uracil synthesis by photodehydrogenation". Nature. 263 (5575): 350–351. Bibcode:1976Natur.263..350C. doi:10.1038/263350a0. PMID 958495. S2CID 4166393.
- ^ Marlaire R (5 November 2009). "NASA reproduces a building block of life in laboratory". NASA. Archived from the original on 4 March 2016. Retrieved 5 March 2015.
- ^ Nuevo, Michel; Materese, Christopher K.; Sandford, Scott A. (2014). "The Photochemistry of Pyrimidine in Realistic Astrophysical ICES and the Production of Nucleobases". The Astrophysical Journal. 793 (2): 125. Bibcode:2014ApJ...793..125N. doi:10.1088/0004-637x/793/2/125. ISSN 1538-4357. S2CID 54189201.
- ^ Marlaire R (3 Mar 2015). "NASA Ames reproduces the building blocks of life in laboratory". NASA. Archived from the original on 5 March 2015. Retrieved 5 Mar 2015.
- ^ a b c Kochetkov NK, Budovskii EI, eds. (1972). Organic Chemistry of Nucleic Acids. Vol. Part B. New York: Plenum Press. doi:10.1007/978-1-4684-2973-2. ISBN 9781468429756.
- ^ a b c Brown EG (1998). Brown EG (ed.). Ring Nitrogen and Key Biomolecules: The biochemistry of N-heterocycles. Boston, MA: Lluwer Academic Publishers. doi:10.1007/978-94-011-4906-8. ISBN 9780412835704. S2CID 9708198.
- ^ Olson KC, Sun D, Chen G, Sharma AK, Amin S, Ropson IJ, et al. (September 2011). "Characterization of dibenzo[a,l]pyrene-trans-11,12-diol (dibenzo[def,p]chrysene) glucuronidation by UDP-glucuronosyltransferases". Chemical Research in Toxicology. 24 (9): 1549–1559. doi:10.1021/tx200178v. PMC 3177992. PMID 21780761.
- ^ Mazur A, Lichti CF, Prather PL, Zielinska AK, Bratton SM, Gallus-Zawada A, et al. (July 2009). "Characterization of human hepatic and extrahepatic UDP-glucuronosyltransferase enzymes involved in the metabolism of classic cannabinoids". Drug Metabolism and Disposition. 37 (7): 1496–1504. doi:10.1124/dmd.109.026898. PMC 2698943. PMID 19339377.
- ^ De Gregori S, De Gregori M, Ranzani GN, Allegri M, Minella C, Regazzi M (March 2012). "Morphine metabolism, transport and brain disposition". Metabolic Brain Disease. 27 (1): 1–5. doi:10.1007/s11011-011-9274-6. PMC 3276770. PMID 22193538.
- ^ a b Mashiyama ST, Courtemanche C, Elson-Schwab I, Crott J, Lee BL, Ong CN, et al. (July 2004). "Uracil in DNA, determined by an improved assay, is increased when deoxynucleosides are added to folate-deficient cultured human lymphocytes". Analytical Biochemistry. 330 (1): 58–69. doi:10.1016/j.ab.2004.03.065. PMID 15183762.
- ^ Zajac MA, Zakrzewski AG, Kowal MG, Narayan S (2003). "A novel method of caffeine synthesis from uracil". Synthetic Communications. 33 (19): 3291–3297. doi:10.1081/SCC-120023986. S2CID 43220488.
- ^ Ramesh D, Mohanty AK, De A, Vijayakumar BG, Sethumadhavan A, Muthuvel SK, et al. (June 2022). "Uracil derivatives as HIV-1 capsid protein inhibitors: design, in silico, in vitro and cytotoxicity studies". RSC Advances. 12 (27): 17466–17480. Bibcode:2022RSCAd..1217466R. doi:10.1039/D2RA02450K. PMC 9190787. PMID 35765450.
- ^ Ramesh, Deepthi; Vijayakumar, Balaji Gowrivel; Kannan, Tharanikkarasu (2021-05-06). "Advances in Nucleoside and Nucleotide Analogues in Tackling Human Immunodeficiency Virus and Hepatitis Virus Infections". ChemMedChem. 16 (9): 1403–1419. doi:10.1002/cmdc.202000849. ISSN 1860-7179. PMID 33427377. S2CID 231576801.
- ^ Ramesh, Deepthi; Vijayakumar, Balaji Gowrivel; Kannan, Tharanikkarasu (2020-12-01). "Therapeutic potential of uracil and its derivatives in countering pathogenic and physiological disorders". European Journal of Medicinal Chemistry. 207: 112801. doi:10.1016/j.ejmech.2020.112801. ISSN 0223-5234. PMID 32927231. S2CID 221724578.
- ^ Ramesh D, Sarkar D, Joji A, Singh M, Mohanty AK, G Vijayakumar B, et al. (April 2022). "First-in-class pyrido[2,3-d]pyrimidine-2,4(1H,3H)-diones against leishmaniasis and tuberculosis: Rationale, in vitro, ex vivo studies and mechanistic insights". Archiv der Pharmazie. 355 (4): e2100440. doi:10.1002/ardp.202100440. PMID 35106845. S2CID 246474821.
- ^ Hidalgo A, Pompei C, Galli A, Cazzola S (January 2005). "Uracil as an index of lactic acid bacteria contamination of tomato products". Journal of Agricultural and Food Chemistry. 53 (2): 349–355. doi:10.1021/jf0486489. PMID 15656671.
- ^ a b Pozharskii AF, Soldatenkov AT, Katritzky AR (1997). Heterocycles in Life and Society: An introduction to heterocyclic chemistry and biochemistry and the role of heterocycles in science, technology, medicine, and agriculture. New York, NY: John Wiley and Sons. ISBN 9780471960348.
- ^ Vijayakumar, Balaji Gowrivel; Ramesh, Deepthi; Manikandan, K. Santhosh; Theresa, Mary; Sethumadhavan, Aiswarya; Priyadarisini, V. Brindha; Radhakrishnan, E. K.; Mani, Maheswaran; Kannan, Tharanikkarasu (2022-06-01). "Chitosan with pendant (E)-5-((4-acetylphenyl)diazenyl)-6-aminouracil groups as synergetic antimicrobial agents". Journal of Materials Chemistry B. 10 (21): 4048–4058. doi:10.1039/D2TB00240J. ISSN 2050-7518. PMID 35507973. S2CID 248526212.
- ^ Séron K, Blondel MO, Haguenauer-Tsapis R, Volland C (March 1999). "Uracil-induced down-regulation of the yeast uracil permease". Journal of Bacteriology. 181 (6): 1793–1800. doi:10.1128/JB.181.6.1793-1800.1999. PMC 93577. PMID 10074071.