The SRC Homology 3 Domain (or SH3 domain) is a small protein domain of about 60 amino acid residues. Initially, SH3 was described as a conserved sequence in the viral adaptor protein v-Crk. This domain is also present in the molecules of phospholipase and several cytoplasmic tyrosine kinases such as Abl and Src.[1][2] It has also been identified in several other protein families such as: PI3 Kinase, Ras GTPase-activating protein, CDC24 and cdc25.[3][4][5] SH3 domains are found in proteins of signaling pathways regulating the cytoskeleton, the Ras protein, and the Src kinase and many others. The SH3 proteins interact with adaptor proteins and tyrosine kinases. Interacting with tyrosine kinases, SH3 proteins usually bind far away from the active site. Approximately 300 SH3 domains are found in proteins encoded in the human genome. In addition to that, the SH3 domain was responsible for controlling protein-protein interactions in the signal transduction pathways[6] and regulating the interactions of proteins involved in the cytoplasmic signaling.[7]
| SH3 domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
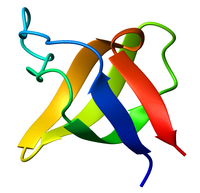 Ribbon diagram of the SH3 domain, alpha spectrin, from chicken (PDB accession code 1SHG), colored from blue (N-terminus) to red (C-terminus). | |||||||||
| Identifiers | |||||||||
| Symbol | SH3_1 | ||||||||
| Pfam | PF00018 | ||||||||
| Pfam clan | CL0010 | ||||||||
| InterPro | IPR001452 | ||||||||
| SMART | SM00326 | ||||||||
| PROSITE | PS50002 | ||||||||
| SCOP2 | 1shf / SCOPe / SUPFAM | ||||||||
| CDD | cd00174 | ||||||||
| |||||||||
Structure edit
The SH3 domain has a characteristic beta-barrel fold that consists of five or six β-strands arranged as two tightly packed anti-parallel β sheets. The linker regions may contain short helices. The SH3-type fold is an ancient fold found in eukaryotes as well as prokaryotes.[8]
Peptide binding edit
The classical SH3 domain is usually found in proteins that interact with other proteins and mediate assembly of specific protein complexes, typically via binding to proline-rich peptides in their respective binding partner. Classical SH3 domains are restricted in humans to intracellular proteins, although the small human MIA family of extracellular proteins also contain a domain with an SH3-like fold.
Many SH3-binding epitopes of proteins have a consensus sequence that can be represented as a regular expression or Short linear motif:
-X-P-p-X-P- 1 2 3 4 5
with 1 and 4 being aliphatic amino acids, 2 and 5 always and 3 sometimes being proline. The sequence binds to the hydrophobic pocket of the SH3 domain. More recently, SH3 domains that bind to a core consensus motif R-x-x-K have been described. Examples are the C-terminal SH3 domains of adaptor proteins like Grb2 and Mona (a.k.a. Gads, Grap2, Grf40, GrpL etc.). Other SH3 binding motifs have emerged and are still emerging in the course of various molecular studies, highlighting the versatility of this domain.
SH3 interactomes edit
SH3 domain-mediated protein-protein interaction networks, i.e., SH3 interactomes, revealed that worm SH3 interactome resembles the analogous yeast network because it is significantly enriched for proteins with roles in endocytosis.[9][10] Nevertheless, orthologous SH3 domain-mediated interactions are highly rewired between worm and yeast.[9]
Proteins with SH3 domain edit
- Signal transducing adaptor proteins
- CDC24
- Cdc25
- PI3 kinase
- Phospholipase
- Ras GTPase-activating protein
- Vav proto-oncogene
- GRB2
- p54 S6 kinase 2 (S6K2)
- SH3D21
- ARMH3 (potentially)
- STAC3
- Some myosins
- SH3 and multiple ankyrin repeat domains: SHANK1, SHANK2, SHANK3
- YAP1
- ARHGAP12
- vexin (VXN)
- TANGO1
- Integrase
- Focal Adhesion Kinase (FAK, PTK2)
- Proline-rich tyrosine kinase (Pyk2, CADTK, PTK2beta)
- TRIP10 (cip4)
See also edit
References edit
- ^ Pawson T, Schlessingert J (July 1993). "SH2 and SH3 domains". Current Biology. 3 (7): 434–42. doi:10.1016/0960-9822(93)90350-W. PMID 15335710. S2CID 53273571.
- ^ Mayer BJ (April 2001). "SH3 domains: complexity in moderation". Journal of Cell Science. 114 (Pt 7): 1253–63. doi:10.1242/jcs.114.7.1253. PMID 11256992.
- ^ Musacchio A, Gibson T, Lehto VP, Saraste M (July 1992). "SH3--an abundant protein domain in search of a function". FEBS Letters. 307 (1): 55–61. doi:10.1016/0014-5793(92)80901-R. PMID 1639195. S2CID 8564342.
- ^ Mayer BJ, Baltimore D (January 1993). "Signalling through SH2 and SH3 domains". Trends in Cell Biology. 3 (1): 8–13. doi:10.1016/0962-8924(93)90194-6. PMID 14731533.
- ^ Pawson T (February 1995). "Protein modules and signalling networks". Nature. 373 (6515): 573–80. doi:10.1038/373573a0. PMID 7531822. S2CID 4324726.
- ^ Schlessinger J (February 1994). "SH2/SH3 signaling proteins". Current Opinion in Genetics & Development. 4 (1): 25–30. doi:10.1016/0959-437X(94)90087-6. PMID 8193536.
- ^ Koch CA, Anderson D, Moran MF, Ellis C, Pawson T (May 1991). "SH2 and SH3 domains: elements that control interactions of cytoplasmic signaling proteins". Science. 252 (5006): 668–74. doi:10.1126/science.1708916. PMID 1708916.
- ^ Whisstock JC, Lesk AM (April 1999). "SH3 domains in prokaryotes". Trends in Biochemical Sciences. 24 (4): 132–3. doi:10.1016/s0968-0004(99)01366-3. PMID 10322416.
- ^ a b Xin, Xiaofeng; Gfeller, David; Cheng, Jackie; Tonikian, Raffi; Sun, Lin; Guo, Ailan; Lopez, Lianet; Pavlenco, Alevtina; Akintobi, Adenrele (2013-01-01). "SH3 interactome conserves general function over specific form". Molecular Systems Biology. 9: 652. doi:10.1038/msb.2013.9. ISSN 1744-4292. PMC 3658277. PMID 23549480.
- ^ Tonikian, Raffi; Xin, Xiaofeng; Toret, Christopher P.; Gfeller, David; Landgraf, Christiane; Panni, Simona; Paoluzi, Serena; Castagnoli, Luisa; Currell, Bridget (2009-10-01). "Bayesian modeling of the yeast SH3 domain interactome predicts spatiotemporal dynamics of endocytosis proteins". PLOS Biology. 7 (10): e1000218. doi:10.1371/journal.pbio.1000218. ISSN 1545-7885. PMC 2756588. PMID 19841731.
External links edit
- Eukaryotic Linear Motif resource motif class LIG_SH3_1
- Eukaryotic Linear Motif resource motif class LIG_SH3_2
- Eukaryotic Linear Motif resource motif class LIG_SH3_3
- Eukaryotic Linear Motif resource motif class LIG_SH3_4
- Eukaryotic Linear Motif resource motif class LIG_SH3_5
- Eukaryotic Linear Motif resource motif class TRG_PEX_1
- Nash Lab Protein Interaction Domains in Signal Transduction - The SH3 domain
- GENEART - Screen your protein against all human SH3 domains in a single phage display cycle