Retroviral integrase (IN) is an enzyme produced by a retrovirus (such as HIV) that integrates (forms covalent links between) its genetic information into that of the host cell it infects.[1] Retroviral INs are not to be confused with phage integrases (recombinases) used in biotechnology, such as λ phage integrase, as discussed in site-specific recombination.
| Integrase Zinc binding domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 solution structure of the n-terminal zn binding domain of hiv-1 integrase (e form), nmr, 38 structures | |||||||||
| Identifiers | |||||||||
| Symbol | Integrase_Zn | ||||||||
| Pfam | PF02022 | ||||||||
| InterPro | IPR003308 | ||||||||
| SCOP2 | 1wjb / SCOPe / SUPFAM | ||||||||
| |||||||||
| Integrase core domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
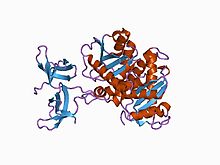 Crystal structure of the RSV two-domain integrase. | |||||||||
| Identifiers | |||||||||
| Symbol | rve | ||||||||
| Pfam | PF00665 | ||||||||
| Pfam clan | CL0219 | ||||||||
| InterPro | IPR001584 | ||||||||
| SCOP2 | 2itg / SCOPe / SUPFAM | ||||||||
| |||||||||
| Integrase DNA binding domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 Crystal structure of the RSV two-domain integrase. | |||||||||
| Identifiers | |||||||||
| Symbol | IN_DBD_C | ||||||||
| Pfam | PF00552 | ||||||||
| InterPro | IPR001037 | ||||||||
| SCOP2 | 1ihw / SCOPe / SUPFAM | ||||||||
| |||||||||
The macromolecular complex of an IN macromolecule bound to the ends of the viral DNA ends has been referred to as the intasome; IN is a key component in this and the retroviral pre-integration complex.[2]
Structure edit
All retroviral IN proteins contain three canonical domains, connected by flexible linkers:[3][4]
- an N-terminal HH-CC zinc-binding domain (a three-helical bundle stabilized by coordination of a Zn(II) cation),
- a catalytic core domain (RNaseH fold),
- a C-terminal DNA-binding domain (SH3 fold).
Crystal and NMR structures of the individual domains and 2-domain constructs of integrases from HIV-1, HIV-2, SIV, and Rous Sarcoma Virus (RSV) have been reported, with the first structures determined in 1994.[5][6] Biochemical data and structural data suggest that retroviral IN functions as a tetramer (dimer-of-dimers), with all three domains being important for multimerization and viral DNA binding.[7] In addition, several host cellular proteins have been shown to interact with IN to facilitate the integration process: e.g., the host factor, human chromatin-associated protein LEDGF, tightly binds HIV IN and directs the HIV pre-integration complex towards highly expressed genes for integration.[8]
Human foamy virus (HFV), an agent harmless to humans, has an integrase similar to HIV IN and is therefore a model of HIV IN function; a 2010 crystal structure of the HFV integrase assembled on viral DNA ends has been determined.[6]
Function and mechanism edit
Integration occurs following production of the double-stranded linear viral DNA by the viral RNA/DNA-dependent DNA polymerase reverse transcriptase.[9]
The main function of IN is to insert the viral DNA into the host chromosomal DNA, an essential step for HIV replication. Integration is a "point of no return" for the cell, which becomes a permanent carrier of the viral genome (provirus). Integration is in part responsible for the persistence of retroviral infections.[10] After integration, the viral gene expression and particle production may take place immediately or at some point in the future, the timing depends on the activity of the chromosomal locus hosting the provirus.[4]
Retroviral INs catalyzes two reactions:[4]
- 3'-processing, in which two or three nucleotides are removed from one or both 3' ends of the viral DNA to expose an invariant CA dinucleotide.
- the strand transfer reaction, in which the processed 3' ends of the viral DNA are covalently ligated to host chromosomal DNA.
Both reactions are catalyzed in the same active site, and involve transesterification without involving a covalent protein-DNA intermediate (in contrast to Ser/Tyr recombinase-catalyzed reactions.[4]
In HIV edit
HIV integrase is a 32kDa viral protein consisting of three domains- N-terminus, catalytic core domain, and C-terminus, which each have distinct properties and functions contributing to the efficacy of HIV integrase.[3]
The N-terminus is composed of 50 amino acid residues which contain a conserved histidine, histidine, cytosine, cytosine sequence which chelates zinc ions, furthermore enhancing the enzymatic activity of the catalytic core domain.[3] As metal chelation is vital in integrase efficacy, it is a target for the development of retroviral therapies.[3]
The catalytic core domain, like the N-terminus, contains highly conserved amino acid residues -Asp64, Asp116, Glu152- as the conserved DDE (Asp-Asp-Glu) motif contributes to the endonuclease and polynucleotide transferase functions of integrase. Mutations in these regions inactivates integrase and prevents genome integration.[3]
The C-terminus domain binds to host DNA non-specifically and stabilizes the integration complex.[3]
Integration mechanism edit
Following synthesis of HIV's doubled stranded DNA genome, integrase binds to the long tandem repeats flanking the genome on both ends. Using its endonucleolytic activity, integrase cleaves a di or trinucleotide from both 3' ends of the genome in a processing known as 3'-processing.[11] The specificity of cleavage is improved through the use of cofactors such as Mn2+ and Mg2+ which interact with the DDE motif of the catalytic core domain, acting as cofactors to integrase function.[11]
The newly generated 3'OH groups disrupt the host DNA's phosphodiester linkages through SN2-type nucleophilic attack.[6] The 3' ends are covalently linked to the target DNA. The 5' over hangs of the viral genome are then cleaved using host repair enzymes, those same enzymes are believed to be responsible for the integration of the 5' end into the host genome forming the provirus.[6][11]
Antiretroviral therapy edit
In November 2005, data from a phase 2 study of an investigational HIV integrase inhibitor, MK-0518, demonstrated that the compound has potent antiviral activity. On October 12, 2007, the Food and Drug Administration (U.S.) approved the integrase inhibitor Raltegravir (MK-0518, brand name Isentress). The second integrase inhibitor, elvitegravir, was approved in the U.S. in August 2012.
See also edit
References edit
- ^ Beck BJ, Freudenreich O, Worth JL (2010). "Patients with Human Immunodeficiency Virus Infection and Acquired Immunodeficiency Syndrome". Massachusetts General Hospital Handbook of General Hospital Psychiatry. Elsevier. pp. 353–370. doi:10.1016/b978-1-4377-1927-7.00026-1. ISBN 9781437719277.
- ^ Masuda T (2011). "Non-Enzymatic Functions of Retroviral Integrase: The Next Target for Novel Anti-HIV Drug Development". Frontiers in Microbiology. 2: 210. doi:10.3389/fmicb.2011.00210. PMC 3192317. PMID 22016749.
- ^ a b c d e f Jóźwik IK, Passos DO, Lyumkis D (September 2020). "Structural Biology of HIV Integrase Strand Transfer Inhibitors". Trends in Pharmacological Sciences. 41 (9): 611–626. doi:10.1016/j.tips.2020.06.003. PMC 7429322. PMID 32624197.
- ^ a b c d Delelis O, Carayon K, Saïb A, Deprez E, Mouscadet JF (December 2008). "Integrase and integration: biochemical activities of HIV-1 integrase". Retrovirology. 5 (1): 114. doi:10.1186/1742-4690-5-114. PMC 2615046. PMID 19091057.
- ^ Lodi PJ, Ernst JA, Kuszewski J, Hickman AB, Engelman A, Craigie R, et al. (August 1995). "Solution structure of the DNA binding domain of HIV-1 integrase". Biochemistry. 34 (31): 9826–9833. doi:10.1021/bi00031a002. PMID 7632683.
- ^ a b c d Choi E, Mallareddy JR, Lu D, Kolluru S (October 2018). "Recent advances in the discovery of small-molecule inhibitors of HIV-1 integrase". Future Science OA. 4 (9): FSO338. doi:10.4155/fsoa-2018-0060. PMC 6222271. PMID 30416746.
- ^ Hare S, Di Nunzio F, Labeja A, Wang J, Engelman A, Cherepanov P (July 2009). Luban J (ed.). "Structural basis for functional tetramerization of lentiviral integrase". PLOS Pathogens. 5 (7): e1000515. doi:10.1371/journal.ppat.1000515. PMC 2705190. PMID 19609359.
- ^ Craigie R, Bushman FD (July 2012). "HIV DNA integration". Cold Spring Harbor Perspectives in Medicine. 2 (7): a006890. doi:10.1101/cshperspect.a006890. PMC 3385939. PMID 22762018.
- ^ Burdick RC, Pathak VK (January 2021). "Efficient HIV-1 in vitro reverse transcription: optimal capsid stability is required". Signal Transduction and Targeted Therapy. 6 (1): 13. doi:10.1038/s41392-020-00458-3. PMC 7804106. PMID 33436564.
- ^ Maldarelli F (February 2016). "The role of HIV integration in viral persistence: no more whistling past the proviral graveyard". The Journal of Clinical Investigation. 126 (2): 438–447. doi:10.1172/JCI80564. PMC 4731194. PMID 26829624.
- ^ a b c Mahboubi-Rabbani M, Abbasi M, Hajimahdi Z, Zarghi A (2021). "HIV-1 Reverse Transcriptase/Integrase Dual Inhibitors: A Review of Recent Advances and Structure-activity Relationship Studies". Iranian Journal of Pharmaceutical Research. 20 (2): 333–369. doi:10.22037/ijpr.2021.115446.15370. PMC 8457747. PMID 34567166.
Further reading edit
- Maertens GN, Engelman AN, Cherepanov P (January 2022). "Structure and function of retroviral integrase". Nature Reviews. Microbiology. 20 (1): 20–34. doi:10.1038/s41579-021-00586-9. PMC 8671357. PMID 34244677. S2CID 235787691.
External links edit
- PDB-101 Molecule of the Month: 135 HIV Integrase
- Integrases at the U.S. National Library of Medicine Medical Subject Headings (MeSH)