In situ hybridization (ISH) is a type of hybridization that uses a labeled complementary DNA, RNA or modified nucleic acid strand (i.e., a probe) to localize a specific DNA or RNA sequence in a portion or section of tissue (in situ) or if the tissue is small enough (e.g., plant seeds, Drosophila embryos), in the entire tissue (whole mount ISH), in cells, and in circulating tumor cells (CTCs). This is distinct from immunohistochemistry, which usually localizes proteins in tissue sections.


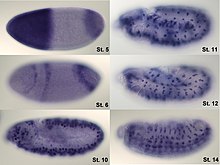

In situ hybridization is used to reveal the location of specific nucleic acid sequences on chromosomes or in tissues, a crucial step for understanding the organization, regulation, and function of genes. The key techniques currently in use include in situ hybridization to mRNA with oligonucleotide and RNA probes (both radio-labeled and hapten-labeled), analysis with light and electron microscopes, whole mount in situ hybridization, double detection of RNAs and RNA plus protein, and fluorescent in situ hybridization to detect chromosomal sequences. DNA ISH can be used to determine the structure of chromosomes. Fluorescent DNA ISH (FISH) can, for example, be used in medical diagnostics to assess chromosomal integrity. RNA ISH (RNA in situ hybridization) is used to measure and localize RNAs (mRNAs, lncRNAs, and miRNAs) within tissue sections, cells, whole mounts, and circulating tumor cells (CTCs). In situ hybridization was invented by American biologists Mary-Lou Pardue and Joseph G. Gall.[1][2][3]
Challenges of in-situ hybridization
editIn situ hybridization is a powerful technique for identifying specific mRNA species within individual cells in tissue sections, providing insights into physiological processes and disease pathogenesis. However, in situ hybridization requires that many steps be taken with precise optimization for each tissue examined and for each probe used. In order to preserve the target mRNA within tissues, it is often required that crosslinking fixatives (such as formaldehyde) be used.[4]
In addition, in-situ hybridization on tissue sections require that tissue slices be very thin, usually 3 μm to 7 μm in thickness. Common methods of preparing tissue sections for in-situ hybridization processing include cutting specimens with a cryostat or a Compresstome tissue slicer. A cryostat takes fresh or fixed tissue and immerses it into liquid nitrogen for flash freezing. Then tissue is embedded in freeze media called OCT and thin sections are cut. Obstacles include getting freeze artifacts on tissue that may interfere with proper mRNA staining. The Compresstome cuts tissue into thin slices without a freeze process; free-floating sections are cut after being embedded in agarose for stability. This method avoids freezing tissue and thus associated freeze artifacts. The process is permanent and irreversible once its complete.[5]
Process
editFor hybridization histochemistry, sample cells and tissues are usually treated to fix the target transcripts in place and to increase access of the probe. As noted above, the probe is either a labeled complementary DNA or, now most commonly, a complementary RNA (riboprobe). The probe hybridizes to the target sequence at elevated temperature, and then the excess probe is washed away (after prior hydrolysis using RNase in the case of unhybridized, excess RNA probe). Solution parameters such as temperature, salt, and/or detergent concentration can be manipulated to remove any non-identical interactions (i.e., only exact sequence matches will remain bound). Then, the probe that was labeled with either radio-, fluorescent- or antigen-labeled bases (e.g., digoxigenin) is localized and quantified in the tissue using either autoradiography, fluorescence microscopy, or immunohistochemistry, respectively. ISH can also use two or more probes, labeled with radioactivity or the other non-radioactive labels, to simultaneously detect two or more transcripts.
An alternative technology, branched DNA assay, can be used for RNA (mRNA, lncRNA, and miRNA ) in situ hybridization assays with single molecule sensitivity without the use of radioactivity. This approach (e.g., ViewRNA assays) can be used to visualize up to four targets in one assay, and it uses patented probe design and bDNA signal amplification to generate sensitive and specific signals. Samples (cells, tissues, and CTCs) are fixed, then treated to allow RNA target accessibility (RNA un-masking). Target-specific probes hybridize to each target RNA. Subsequent signal amplification is predicated on specific hybridization of adjacent probes (individual oligonucleotides [oligos] that bind side by side on RNA targets). A typical target-specific probe will contain 40 oligonucleotides, resulting in 20 oligo pairs that bind side-by-side on the target for detection of mRNA and lncRNA, and 2 oligos or a single pair for miRNA detection. Signal amplification is achieved via a series of sequential hybridization steps. A pre-amplifier molecule hybridizes to each oligo pair on the target-specific RNA, then multiple amplifier molecules hybridize to each pre-amplifier. Next, multiple label probe oligonucleotides (conjugated to alkaline phosphatase or directly to fluorophores) hybridize to each amplifier molecule. A fully assembled signal amplification structure “Tree” has 400 binding sites for the label probes. When all target-specific probes bind to the target mRNA transcript, an 8,000 fold signal amplification occurs for that one transcript. Separate but compatible signal amplification systems enable the multiplex assays. The signal can be visualized using a fluorescence or brightfield microscope.
Basic steps for digoxigenin-labeled probes
edit- permeabilization of cells with proteinase K to open cell membranes (around 25 minutes, not needed for tissue sections or some early-stage embryos)
- binding of mRNAs to marked RNA probe (usually overnight)
- antibody-phosphatase binding to RNA-probe (some hours)
- staining of antibody (e.g., with alkaline phosphatase)
The protocol takes around 2–3 days and takes some time to set up. Some companies sell robots to automate the process (e.g., CEM InsituPro). As a result, large-scale screenings have been conducted in laboratories on thousands of genes. The results can usually be accessed via websites (see external links).[6]
See also
editReferences
edit- ^ O'Connor, Clare. "Fluorescence In Situ Hybridization (FISH)". Nature Education.
- ^ Gall, JG; Pardue, ML (June 1969). "Formation and detection of RNA-DNA hybrid molecules in cytological preparations". Proceedings of the National Academy of Sciences of the United States of America. 63 (2): 378–83. Bibcode:1969PNAS...63..378G. doi:10.1073/pnas.63.2.378. PMC 223575. PMID 4895535.
- ^ Gall, Joe. "Albert Lasker Award for Special Achievement in Medical Science". Lasker Foundation.
- ^ Lehmann, Ruth; Tautz, Diethard (1994-01-01), Goldstein, Lawrence S. B.; Fyrberg, Eric A. (eds.), Chapter 30 in Situ Hybridization to RNA, Methods in Cell Biology, vol. 44, Academic Press, pp. 575–598, doi:10.1016/s0091-679x(08)60933-4, ISBN 9780125641456, PMID 7535885, retrieved 2021-02-21
- ^ Brown, Lindsay A.; Huntsman, David (2007-05-01). "Fluorescent in situ hybridization on tissue microarrays: challenges and solutions". Journal of Molecular Histology. 38 (2): 151–157. doi:10.1007/s10735-006-9069-y. ISSN 1567-2387. PMID 17216303. S2CID 6363208.
- ^ Panoskaltsis-Mortari, A.; Bucy, R. P. (February 1995). "In situ hybridization with digoxigenin-labeled RNA probes: facts and artifacts". BioTechniques. 18 (2): 300–307. ISSN 0736-6205. PMID 7727134.
- Jin, L; Lloyd, RV (1997). "In situ hybridization: methods and applications". Journal of Clinical Laboratory Analysis. 11 (1): 2–9. doi:10.1002/(SICI)1098-2825(1997)11:1<2::AID-JCLA2>3.0.CO;2-F. PMC 6760707. PMID 9021518.
- Comprehensive and annotated in situ hybridization histochemistry
- RNA sequencing of pancreatic circulating tumour cells implicates WNT signalling in metastasis
- The Local Transcriptome in the Synaptic Neuropil Revealed by Deep Sequencing and High-Resolution Imaging
External links
edit- In+Situ+Hybridization at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- In Situ Hybridization of RNA and miRNA Probes to cells, CTCs, and tissues
- Whole-Mount In Situ Hybridization of RNA Probes to Plant Tissues
- Preparation of Complex DNA Probe Sets for 3D FISH with up to Six Different Fluorochromes
- Transcript In Situ Hybridization of Whole-Mount Embryos for Phenotype Analysis of RNAi-Treated Drosophila
- in-situ databases: